The manta code continues
Genetics are a powerful tool in the conservation arsenal. The Manta Trust is collecting samples from vulnerable manta rays all over the globe, and genetic analyses will help us to correctly identify species and discover which populations are most in trouble.
My earliest memory of the ocean is a little unusual. I was on a family beach holiday in the UK when I was faced by this enormous, deafening wall of water and ran away as fast as my little legs would carry me, crying my eyes out. My parents’ response to this behaviour was to pick me up, and throw me straight in at the deep end (pun intended!) – no messing about, and I can’t thank them enough for it!
Growing up, I read endless books about wildlife, and the ocean, eager to know everything there was to know. Then...



Manta ray genetics project
To use novel genetic tools to better understand, conserve and manage manta rays.
Manta rays play a key role in marine ecosystems and are of great value to humans, therefore it is vital that we use these powerful genetic techniques to better understand manta ray biology and further support their long-term conservation and management.
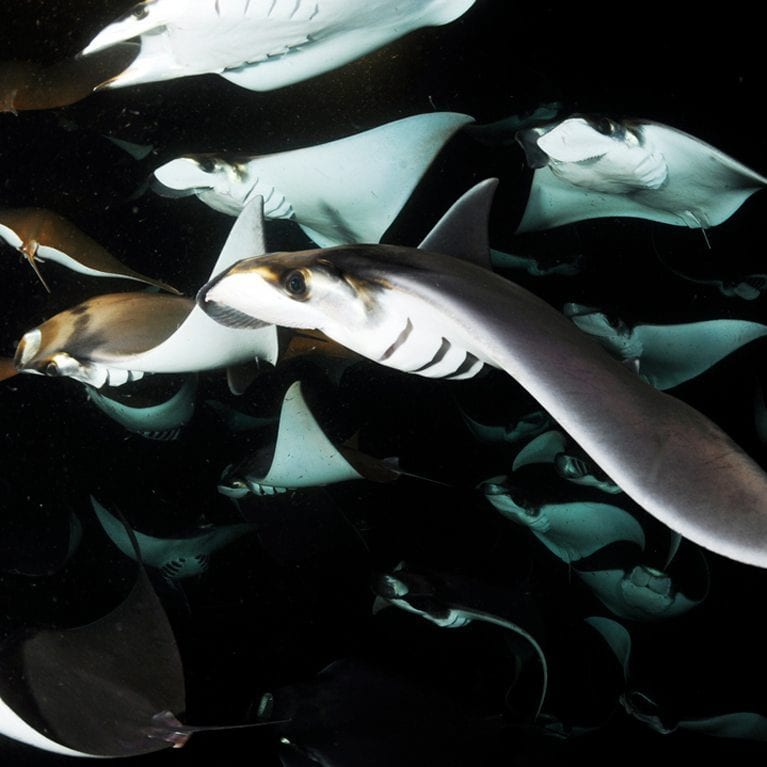
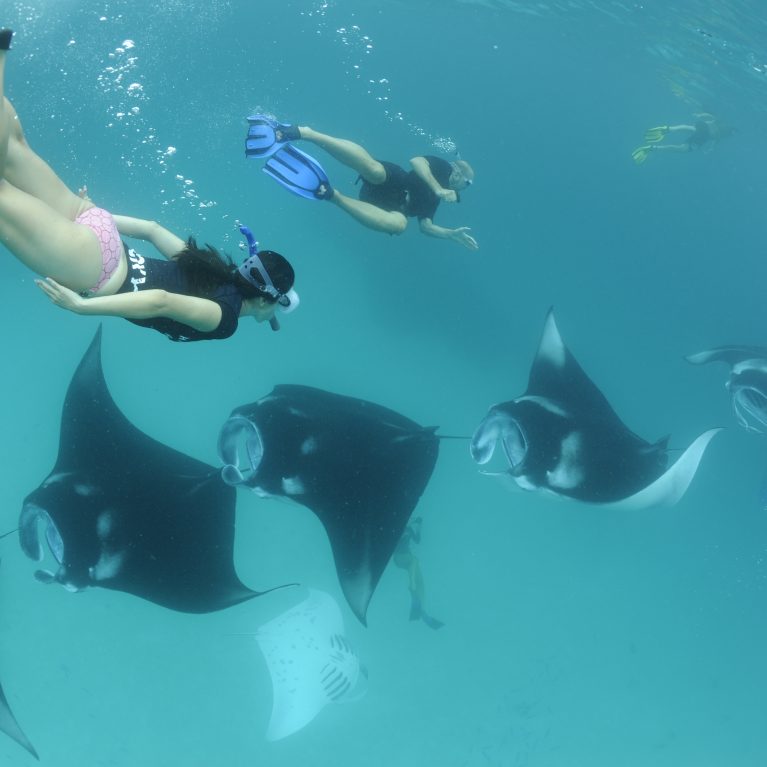
Manta rays are some of the most beautiful and charismatic animals in our oceans and yet we know relatively little about how they came to be here and how they are likely to respond to the increasing threats being put on them.
There are two species of manta rays currently described: Manta alfredi, which leads a resident lifestyle around reefs and atolls, and Manta birostris, which leads a more migratory lifestyle, often crossing international borders. Both species present two colour-morphs – black and chevron – that are found at differing frequencies across the globe. Little is understood about the molecular basis for speciation and morphological variation of manta rays, yet in the face of increasing global fishing pressure and climate change this is vitally important to understand.
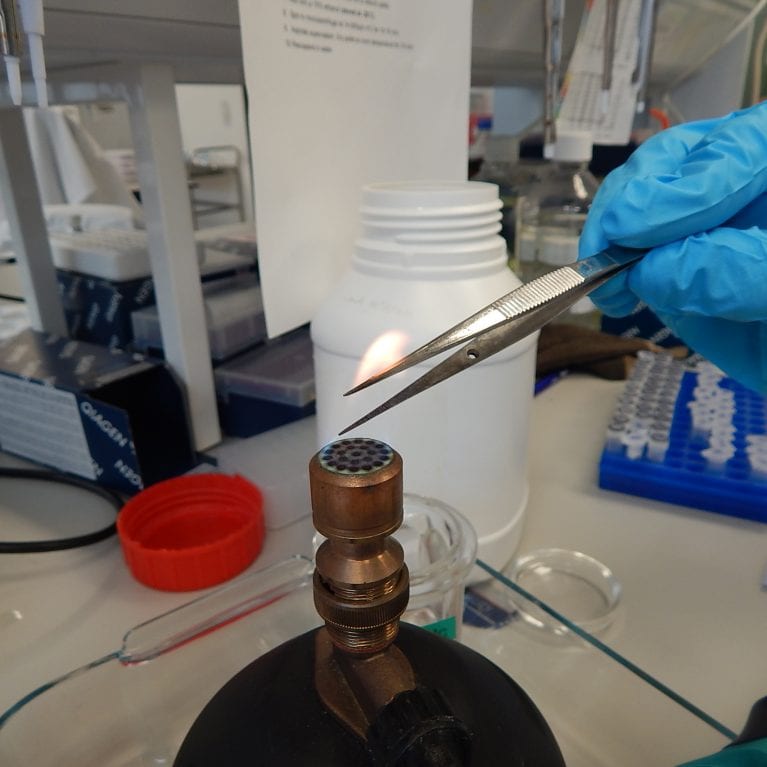
It is now possible to address such questions using next-generation DNA sequencing technologies and bioinformatics. These techniques allow us to sequence and understand more genetic data than ever before, giving us a much better understanding of how species evolve and adapt. In doing so, populations with local genetic adaptations can be identified and subsequently prioritised for conservation due to their increased potential to adapt to environmental change.
This project is one of the first to make the transition to conservation and adaptation genomics in the elasmobranch realm, and it will inevitably help to fill the striking gaps in available elasmobranch sequence data. Not only will this data help to answer questions about manta rays, it will also equip elasmobranch geneticists worldwide with a powerful set of tools for advancing their research goals.
- Work alongside the Manta Trust, which has more than 12 field projects across the globe, to enhance and establish an extensive set of manta ray tissue samples.
- Use comparative genomic methods to identify gene regions showing differentiation between species and populations. Thus, we’ll be able to address how these regions link to differences observed in the wild, for example, in their migratory capabilities.
- Compare the transcriptomes of both colour-morphs to identify which regions are differentially expressed therefore indicating their role in pigmentation patterns. How has this evolved in different populations? Does it confer a selective advantage?

