Lessons in the laboratory
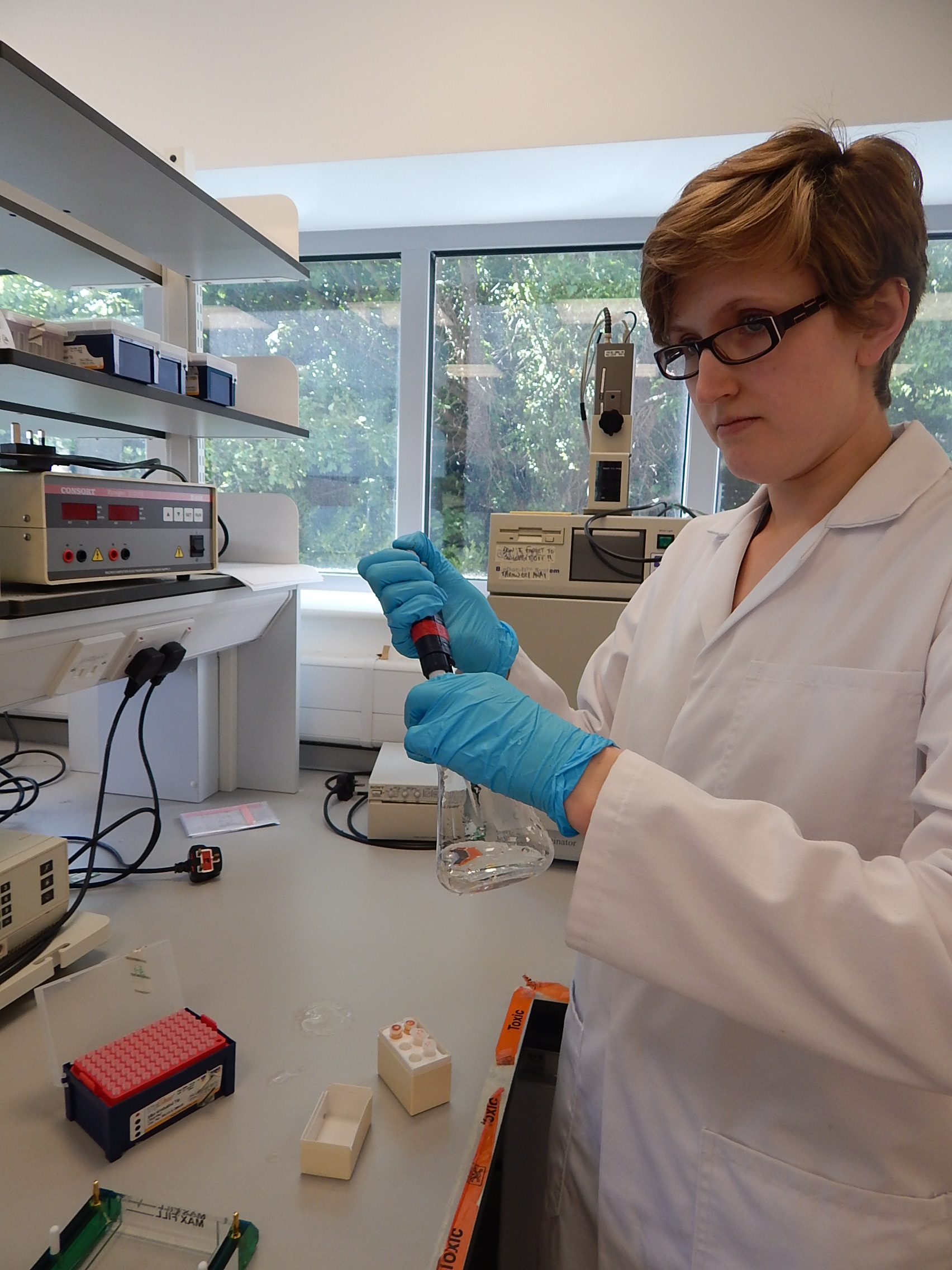
It’s a standard Monday morning when I walk into the lab for the first time since starting my PhD on the genetics of manta and mobula rays. Machines are whirring away around me and all the other PhD students and researchers are super busy with their own projects.
My project aims to develop genetic tools that will assist in the conservation of manta and mobula rays – which is a pretty broad endeavour. So where on earth do I begin? Fortunately, thanks to support from the Save Our Seas Foundation, colleagues from The Manta Trust were working flat out to collect tissue samples from all over the world before I came on the scene. Lots of other researchers got involved too and the result is that I have plenty of material to work with.
One of the main aims of this project is to produce tools that can identify a manta or a mobula ray, or a body part such as a gill plate, to species level from the genetic information contained within it. This will play a major role in the enforcement of conservation regulations. Later on, work will begin on a similar tool to identify in which region the sample originated, and therefore we’ll be able to know where in the world that specific individual came from. This will show which regions are contributing most to the exploitation of manta and mobula rays for their gill plates, and thus where outreach and enforcement are most needed. All of this can be done with DNA extracted from a small piece of tissue, and since more than 800 tissue samples have been collected and received to date, I have so much to do!

The amount of tissue needed for DNA analysis is really small!. © Photo by Jane Hosegood | © Manta Trust

By making use of genetic tools, scenes like this could become far rarer. © Photo by Daniel Fernando | Manta Trust
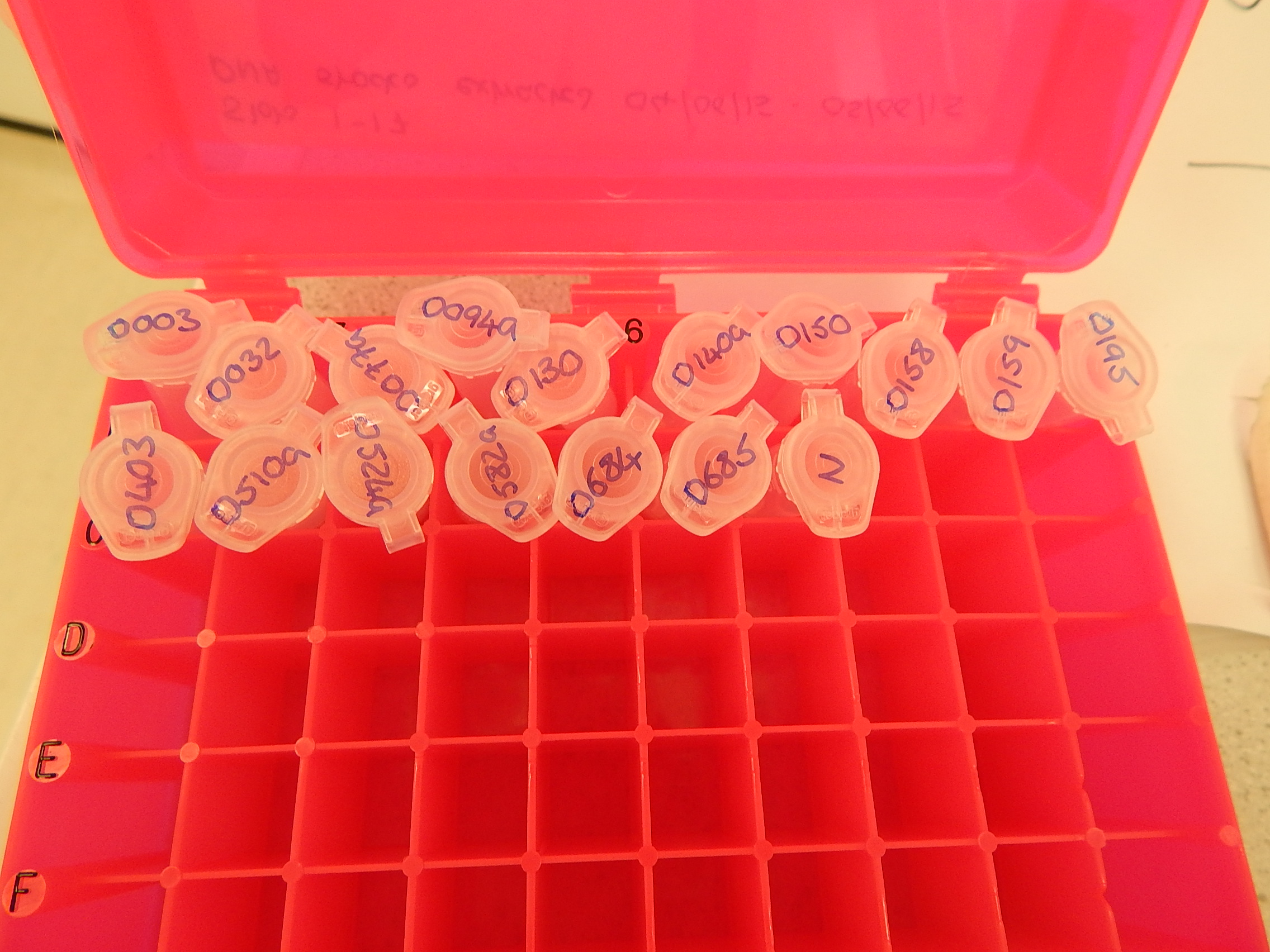
So how do you go from a small piece of tissue to actual DNA that you can work with? And even then, what do you do with it? The extraction process is fairly straightforward and, depending on the technique you use, DNA can be extracted from a batch of samples in a day’s work. In a nutshell, the process involves digesting a tiny piece of tissue with an enzyme to release the DNA from the cells and then removing, or breaking down, all the other bits and pieces that you’re not interested in. The result is a two-millilitre tube with a few drops of clear liquid inside. It’s very easy to confuse different samples, so a lot of meticulous labelling goes hand in hand with any research involving DNA. It’s also very important to consider the possibility of contamination, when DNA from one individual ends up in a tube containing DNA from another individual. If that happens, you have no way of knowing which individual your results apply to, so it can really mess up your analyses.
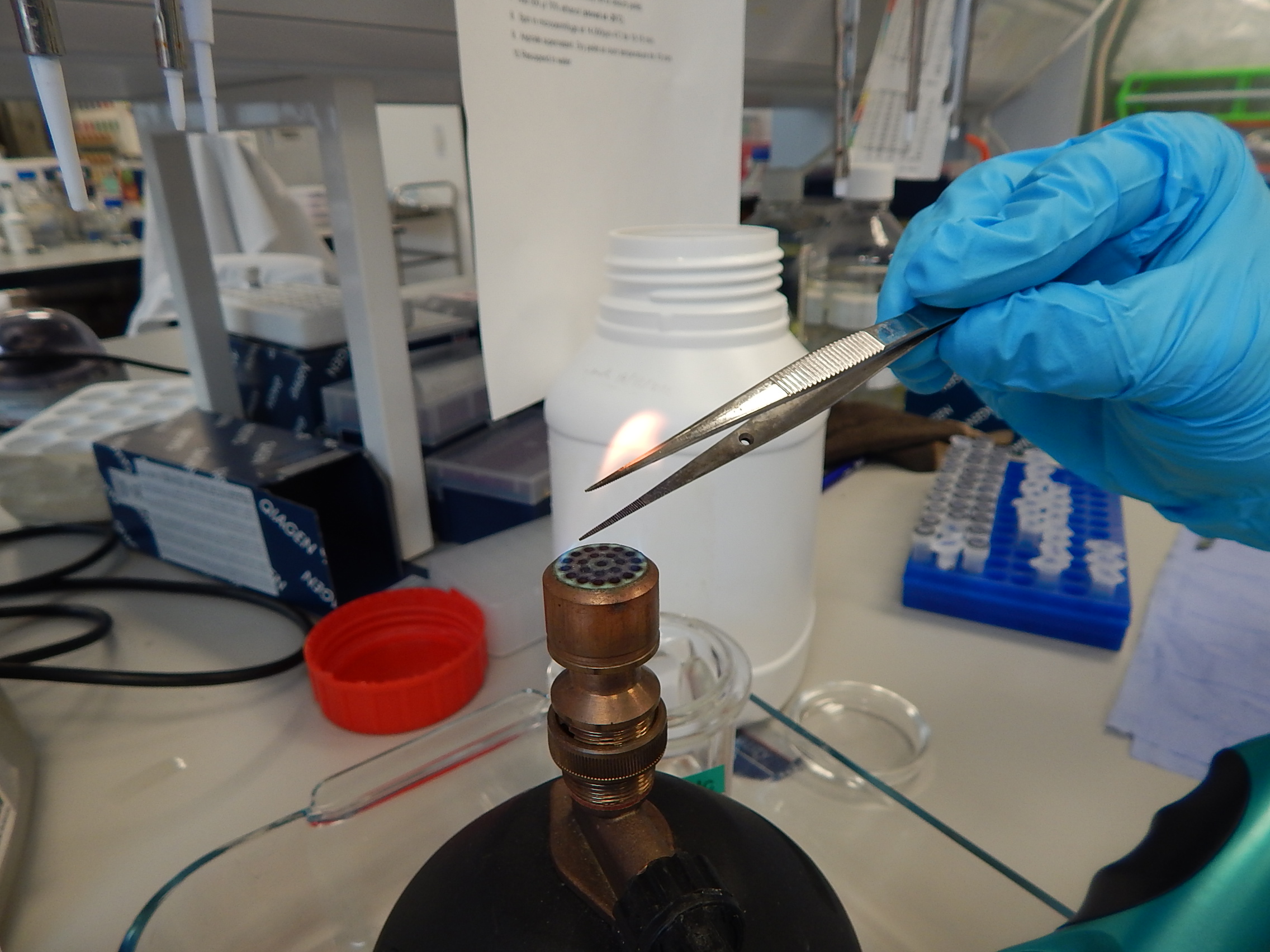

Sterilising tweezers used to handle tissue samples so as not to mix up DNA from different samples. © Photo by Jane Hosegood | Manta Trust
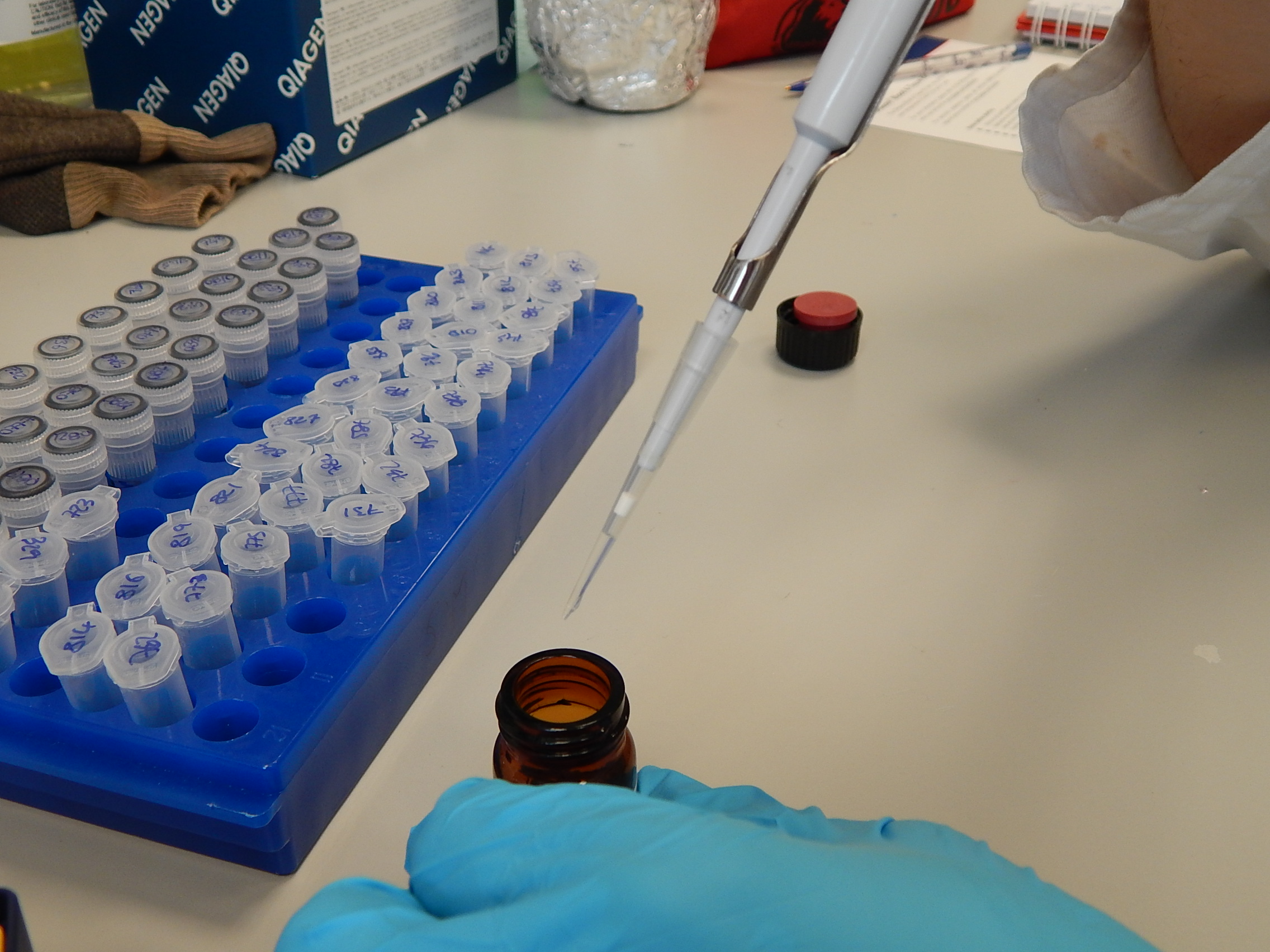
Pipettes have disposable tips to reduce the risk of contamination. © Photo by Jane Hosegood | Manta Trust
So now you have your DNA, but how do you know the extraction has been successful? And that you have good DNA to work with? There are a couple of things you can do: you can measure the amount of DNA based on the wavelength of light reflected back from the extract, or you can use an electric current to run the extract on a special gel, which can be seen under an ultraviolet light.
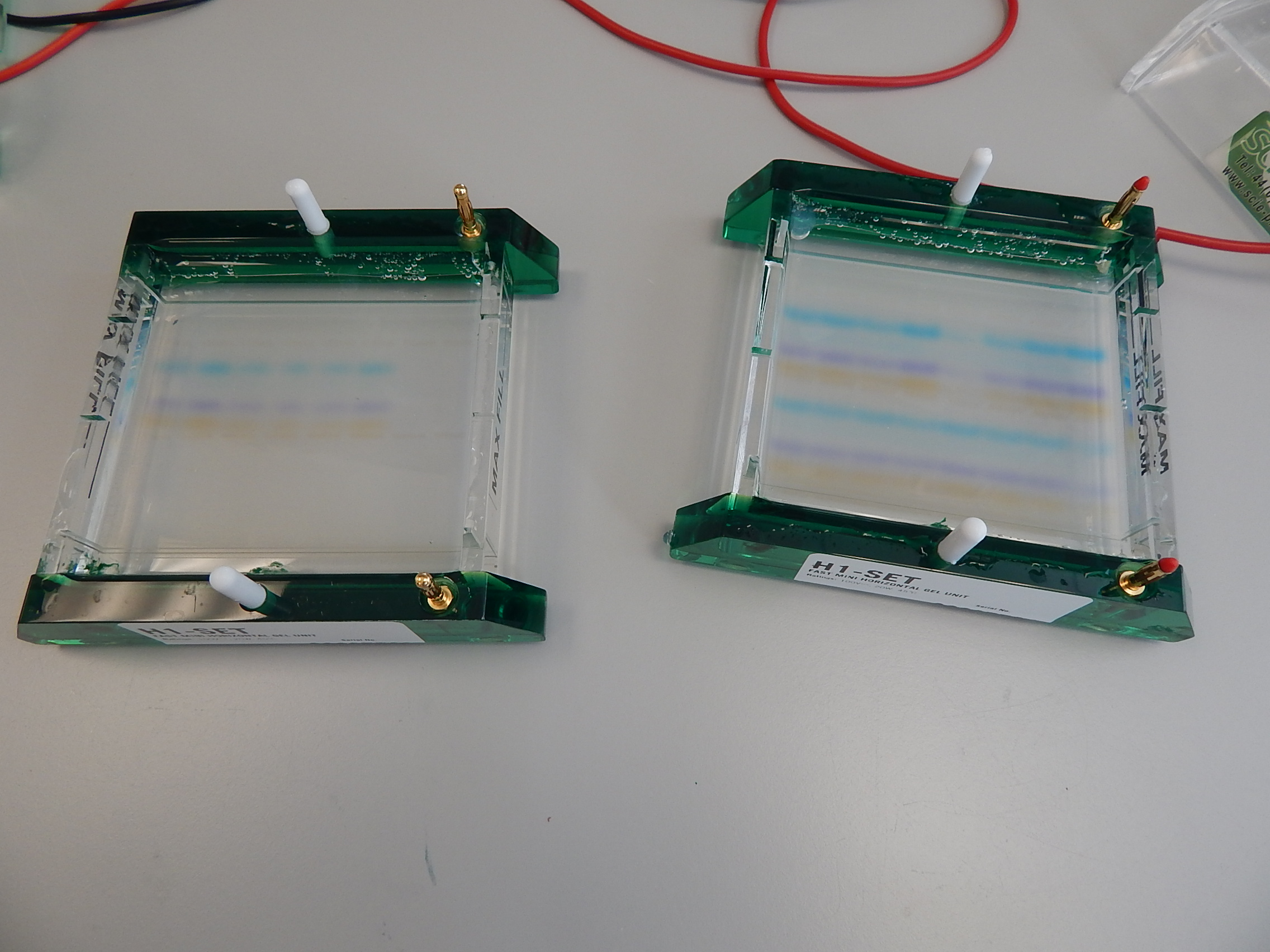

When the gel is run, DNA fragments have moved through at different rates, according to size. © Photo by Jane Hosegood | Manta Trust
Now that you know the extracted DNA is good enough to work with, the options for what you can do with it are virtually endless. I will be using advances in genetic techniques to look for areas of the genome that are consistently different between species, but always the same within a species. It is these areas that can then be specifically targeted and used to prove which species a sample came from – and thus ensure that conservation measures preventing trade in manta parts are effective.
Genetic tools for the conservation of manta and mobula rays are coming – watch this space!