Coding for conservation
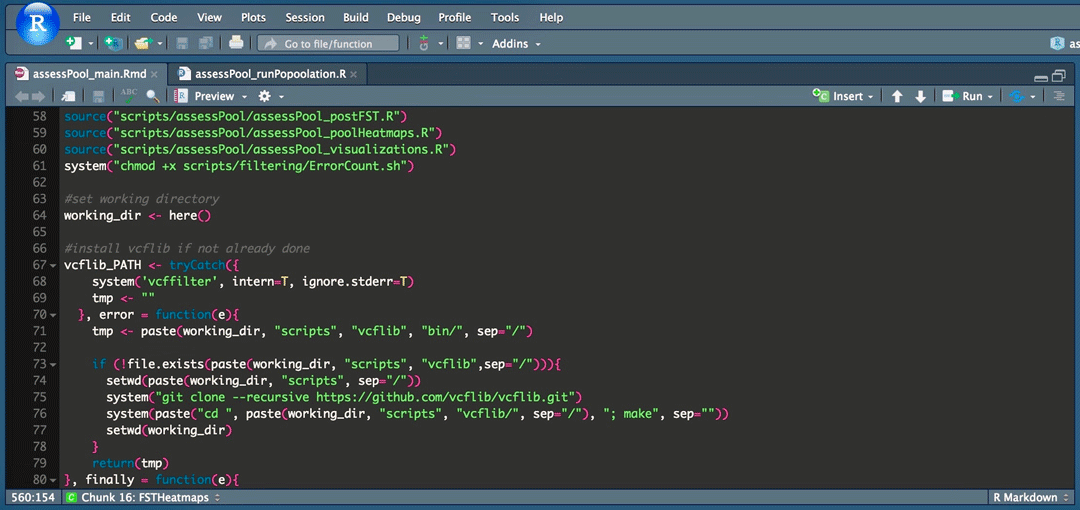
Image © Derek Kraft
>Last login: Sun Aug 24 07:02:05 on ttys000
>Dereks-MacBook-Pro:~ derekkraft$ echo ‘hello welcome to my blog post’
>hello welcome to my blog post
When meeting young hopeful marine biologists, I’m usually introduced as “this is Derek, he studies sharks” which is true and I honestly couldn’t be more proud to be introduced this way. I’ve wanted to study sharks since… well… I was a young hopeful marine biologist. Somewhere during these conversations, some form of the question “What can I do to better prepare myself to be a marine biologist?” usually arises. My answer often surprises them: “Take as many computer science classes as you can.” Then I wait during the inevitable pause as they realize their ears did not deceive them but I did actually suggest taking computer science classes. “Really?” they reply a little confused and deflated, perhaps hoping that I would have instead suggested taking up freediving or immediately booking a one-way ticket to Hawaii. However, I feel that computer science experience will serve them better as they pursue their dream field.
As technology advances, we are getting better and better at collecting large quantities of data. This produces HUGE data sets. Whether it’s tracking sharks with satellite tags, collecting multiple environmental variables over several sites, quantifying photo mosaics of patch reefs to identify coral cover or all of the above all at once, these data sets all require computer programs to assess. Sometimes the ability to collect the data outruns the tools to actually analyze it all. Furthermore, what we want to do with the data changes from project to project, which means that scientists have to customize the data analysis from project to project. This is where a basic knowledge of computer coding is not only helpful but often critical.
When I started graduate school five years ago, (five years already?… oh dear…) I imagined diving with sharks, long days on boats, copious amounts of sunscreen, and all of the other wonderful things scientists tend to post on social media or their Save Our Seas blogs. Two years in, and my dive gear was dusty, no fieldwork in sight, and I was rather pale from all my time spent in the lab preparing DNA from silky sharks. However, all my indoor work had paid off as I had just received my first genomic data set back from the sequencer. Unfortunately, it was in the form of 60gb files of four-letter code that was supposed to tell me something significant about silky sharks’ ecology. That’s when I realized that I completely lacked the computer skills to actually do anything with all the information I had just received. Now what?
I had to start from scratch. In the middle of my second year of grad school, I learned what the command line was (basically a window to talk to your computer in computer language, see Figure 1). There I was, staring at the blinking green cursor, which was waiting impatiently for me to type something meaningful. I had no idea what to do.
Figure 1. A command line window. Image © Derek Kraft
The learning curve was steep. I had to learn the difference between file, directory, working directory, script, executable file, figure out the exact formatting (known as syntax), and what the heck is this error about my PATH? This was just scratching the surface. Every move you make in the command line requires a very specific input, whether it’s copying a file (cp file_name), moving a file (mv filename where_file_goes), or on the more advanced side, performing a genomic assembly (spades.py –dataset libraries.yaml -k 81,91,99,121 –careful -t 30 -m 300 -o ./spades_output). Every command needs the correct input files in the correct format, so I often found myself sorting through various error outputs trying to find the problem. Eventually, I started googling the errors, as I was certainly not the first person to have issues with this code. Sometimes I found the solution quickly, other times it would take me a day or two. However, just like learning any new language, the more I used it the easier it became.
During my third year of graduate school, I took a statistics class using R (highly recommended) which is a different coding language than the command line uses. Running R in a program called Rstudio is more user-friendly because spaces don’t matter as much, it shows you the objects you have created in another window, and visualizes outputs nicely, tools the command line does not provide. R is great for running statistical analysis, visualizing data, and I do enjoy it for making maps. However, I still find it easier to manipulate data sets in the command line and it runs faster when performing high processing jobs. Therefore it’s helpful to have multiple tools in your computer programming toolbox.
I really had no idea that computer coding was going to be a part of my marine biology education. Given that I had zero experience with this sort of thing before I started, I felt very unprepared to start my data analysis and it was very overwhelming. This struggle still feels very real; even after a few years of working in these languages, I still have so much to learn. However, as I make progress, become faster, and start to feel confident, coding does start to feel rewarding, and maybe even a little fun. It’s like solving some sort of odd puzzle where the outcome is yet another file. But, this time it’s in the correct format and is ready for the next step in the analysis (where I’m bound to hit more errors, but it’s progress nonetheless
I would highly recommend checking out the book Practical Computer For Biologists by Steven H. D. Haddock and Casey W. Dunn. It lays out the basics in an easy-to-follow format while leading you through examples that you can perform yourself to get the practice you need, sort of like actually speaking Spanish in Spanish class. Additionally, I feel most schools are now offering classes in computer coding for biologists, or at least statistics using R. These courses are a great way to get started and get you coding! Plus you then make contacts with professors who can help you with your future bioinformatic endeavours.

Figure 2. My daily work station. Photo © Derek Kraft
Although I’m still not diving as often as I’d like, I don’t need as much sunscreen as I’d hoped, and I spend more time with command lines than I do with anchor lines, the computer programming skills I’ve developed are crucial work as marine biology who studies shark! Currently I’m using a combination of R and command-line scripts to explore silky shark genomes (Figure 2, which is how I spend most of my days in graduate school), in search of regional diagnostic genes so I can look at shark samples of unknown origin and match them back to the region where they came from. This allows me to take silky shark fins from the shark fin industry and match it back to the global region that it came from. This will allow us to identify harvest hot spots and account for harvest from the fin trade down to a regional basis, or at least by ocean basin, providing much-needed harvest data for managers to better protect his vulnerable species. Check out my project page below for more details!
